About us
Learn how GA4GH helps expand responsible genomic data use to benefit human health.
Learn how GA4GH helps expand responsible genomic data use to benefit human health.
Our Strategic Road Map defines strategies, standards, and policy frameworks to support responsible global use of genomic and related health data.
Discover how a meeting of 50 leaders in genomics and medicine led to an alliance uniting more than 5,000 individuals and organisations to benefit human health.
GA4GH Inc. is a not-for-profit organisation that supports the global GA4GH community.
The GA4GH Council, consisting of the Executive Committee, Strategic Leadership Committee, and Product Steering Committee, guides our collaborative, globe-spanning alliance.
The Funders Forum brings together organisations that offer both financial support and strategic guidance.
The EDI Advisory Group responds to issues raised in the GA4GH community, finding equitable, inclusive ways to build products that benefit diverse groups.
Distributed across a number of Host Institutions, our staff team supports the mission and operations of GA4GH.
Curious who we are? Meet the people and organisations across six continents who make up GA4GH.
More than 500 organisations connected to genomics — in healthcare, research, patient advocacy, industry, and beyond — have signed onto the mission and vision of GA4GH as Organisational Members.
These core Organisational Members are genomic data initiatives that have committed resources to guide GA4GH work and pilot our products.
This subset of Organisational Members whose networks or infrastructure align with GA4GH priorities has made a long-term commitment to engaging with our community.
Local and national organisations assign experts to spend at least 30% of their time building GA4GH products.
Anyone working in genomics and related fields is invited to participate in our inclusive community by creating and using new products.
Wondering what GA4GH does? Learn how we find and overcome challenges to expanding responsible genomic data use for the benefit of human health.
Study Groups define needs. Participants survey the landscape of the genomics and health community and determine whether GA4GH can help.
Work Streams create products. Community members join together to develop technical standards, policy frameworks, and policy tools that overcome hurdles to international genomic data use.
GIF solves problems. Organisations in the forum pilot GA4GH products in real-world situations. Along the way, they troubleshoot products, suggest updates, and flag additional needs.
GIF Projects are community-led initiatives that put GA4GH products into practice in real-world scenarios.
The GIF AMA programme produces events and resources to address implementation questions and challenges.
NIF finds challenges and opportunities in genomics at a global scale. National programmes meet to share best practices, avoid incompatabilities, and help translate genomics into benefits for human health.
Communities of Interest find challenges and opportunities in areas such as rare disease, cancer, and infectious disease. Participants pinpoint real-world problems that would benefit from broad data use.
The Technical Alignment Subcommittee (TASC) supports harmonisation, interoperability, and technical alignment across GA4GH products.
Find out what’s happening with up to the minute meeting schedules for the GA4GH community.
See all our products — always free and open-source. Do you work on cloud genomics, data discovery, user access, data security or regulatory policy and ethics? Need to represent genomic, phenotypic, or clinical data? We’ve got a solution for you.
All GA4GH standards, frameworks, and tools follow the Product Development and Approval Process before being officially adopted.
Learn how other organisations have implemented GA4GH products to solve real-world problems.
Help us transform the future of genomic data use! See how GA4GH can benefit you — whether you’re using our products, writing our standards, subscribing to a newsletter, or more.
Join our community! Explore opportunities to participate in or lead GA4GH activities.
Help create new global standards and frameworks for responsible genomic data use.
Align your organisation with the GA4GH mission and vision.
Want to advance both your career and responsible genomic data sharing at the same time? See our open leadership opportunities.
Join our international team and help us advance genomic data use for the benefit of human health.
Discover current opportunities to engage with GA4GH. Share feedback on our products, apply for volunteer leadership roles, and contribute your expertise to shape the future of genomic data sharing.
Solve real problems by aligning your organisation with the world’s genomics standards. We offer software dvelopers both customisable and out-of-the-box solutions to help you get started.
Learn more about upcoming GA4GH events. See reports and recordings from our past events.
Speak directly to the global genomics and health community while supporting GA4GH strategy.
Be the first to hear about the latest GA4GH products, upcoming meetings, new initiatives, and more.
Questions? We would love to hear from you.
Read news, stories, and insights from the forefront of genomic and clinical data use.
Publishes regular briefs exploring laws and regulations, including data protection laws, that impact genomic and related health data sharing
Translates findings from studies on public attitudes towards genomic data sharing into short blog posts, with a particular focus on policy implications
Attend an upcoming GA4GH event, or view meeting reports from past events.
See new projects, updates, and calls for support from the Work Streams.
Read academic papers coauthored by GA4GH contributors.
Listen to our podcast OmicsXchange, featuring discussions from leaders in the world of genomics, health, and data sharing.
Check out our videos, then subscribe to our YouTube channel for more content.
View the latest GA4GH updates, Genomics and Health News, Implementation Notes, GDPR Briefs, and more.
22 Oct 2019
The GA4GH Variation Representation (VR) specification, produced by the Genomic Knowledge Standards Work Steam, provides a flexible framework of computational models, schemas, and algorithms to precisely and consistently exchange genetic variation data across communities.

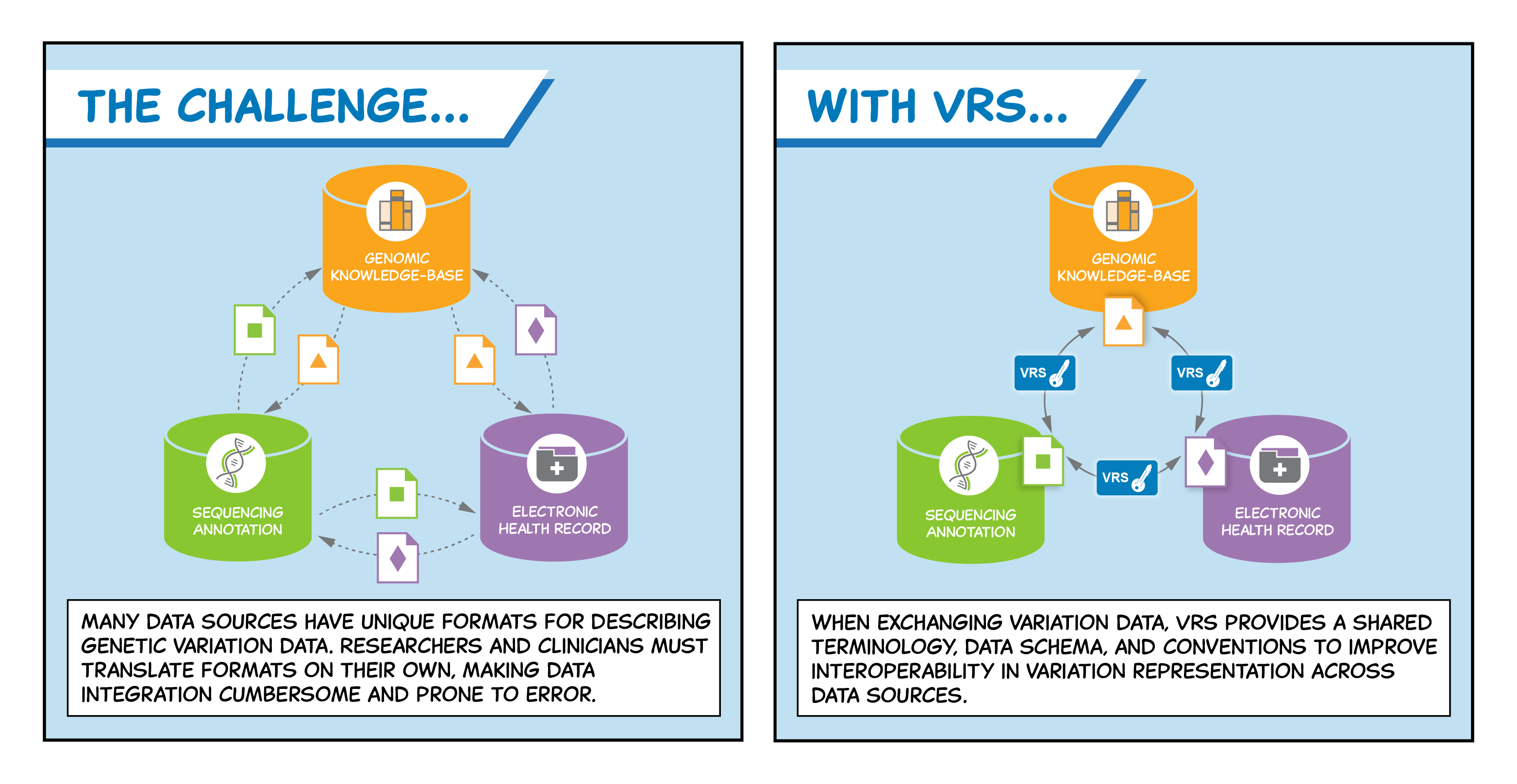
Image Credit: Stephanie Li, GA4GH
Between any two individuals, the human genome differs by around 0.1 percent—meaning each individual has millions of sites in their genome that makes them unique. The impact of genetic variation on human health and disease is complex and immense; in order to accelerate research and advance patient care, it is vital to maximize the value of the world’s genetic variation data and exchange it reliably across diagnostic labs, electronic health records (EHRs), research institutions, and knowledgebases.
The GA4GH Variation Representation (VR) specification, produced by the Genomic Knowledge Standards Work Steam, provides a flexible framework of computational models, schemas, and algorithms to precisely and consistently exchange genetic variation data across communities. The specification, which was developed with input from national information resource providers, major public initiatives, and diagnostic testing laboratories, significantly reduces ambiguity in exchanging variation data. In this way, VR aims to improve the reliability and utility of the clinical annotations that are central to personalized medicine.
“The genomics community lacks a robust standard that enables the unambiguous exchange of genomic data,” said Robert Freimuth, co-lead of the Genomic Knowledge Standards Work Stream and Assistant Professor of Biomedical Informatics in the Division of Digital Health Sciences at the Mayo Clinic. “The VR-Spec is a step toward filling the gap between the exchange mechanisms used by the research, translational, and clinical communities, which is necessary for the implementation of genomic and precision medicine.”
“VR reduces the ambiguity in the way we exchange sequence variation data,” said Reece Hart, a software engineering consultant and principal author of the specification. “By reducing the ambiguity, we improve the precision with which we use genetic variation in research and clinical settings. ”
With common needs across all variant data repositories—such as designating two or more variants as functionally-equivalent—there is a strong need for encoding community-driven best practices in variant representation.
“In practice, each different repository has made its own decisions fairly independently, with no best practices emerging,” said UCSC bioinformatician Melissa Cline, a Driver Project Champion of the BRCA Exchange and an implementer of the VR specification. “The VR specification provides us with a common vocabulary that we can now build upon as a community to define these best practices.”
Alex Wagner, an instructor at Washington University School of Medicine who co-leads the Variant Interpretation for Cancer Consortium, also co-led VR’s development. “We have designed the specification with intent to capture difficult variation concepts, such as structural variation, in a precise, computable way to improve cross-platform search,” said Wagner. “A shared, unified model for these variation forms will be a tremendous asset for normalizing knowledge used in precision oncology.”
The VR specification consists of several key components that together produce a reliable way of describing and transferring genetic variation data:
“Genetic variation is complex. Currently, no single model exists to represent and exchange this data in a consistent, reliable manner,” said product lead Larry Babb, a senior principal software engineer at the Broad Institute of MIT and Harvard. “By addressing this issue, the specification will allow different communities to ‘speak the same language’— whether it’s diagnostic labs and EHR vendors who are collecting samples or investigators who are accessing them.”
The ClinGen Allele Registry, which provides unique variant identifiers for the community, has developed an initial implementation for the VR specification. “Wide adoption of the VR specification will provide a means of collating information about variants [at scale],” according to a statement by Allele Registry authors Aleks Milosavljevic and Ronak Y. Patel of the Baylor College of Medicine. “This capability has so far been lacking and is synergistic with the approach of the Allele Registry and other web services, which support the linking of information on almost one billion distinct variants.”
The VR specification aims to maximize the value of the world’s shared genetic variation data. In addition to building computational models for more complex types of variation, the team has future plans to describe new types of intervals and locations, as well as develop additional concepts for representing state and variation.