About us
Learn how GA4GH helps expand responsible genomic data use to benefit human health.
Learn how GA4GH helps expand responsible genomic data use to benefit human health.
Our Strategic Road Map defines strategies, standards, and policy frameworks to support responsible global use of genomic and related health data.
Discover how a meeting of 50 leaders in genomics and medicine led to an alliance uniting more than 5,000 individuals and organisations to benefit human health.
GA4GH Inc. is a not-for-profit organisation that supports the global GA4GH community.
The GA4GH Council, consisting of the Executive Committee, Strategic Leadership Committee, and Product Steering Committee, guides our collaborative, globe-spanning alliance.
The Funders Forum brings together organisations that offer both financial support and strategic guidance.
The EDI Advisory Group responds to issues raised in the GA4GH community, finding equitable, inclusive ways to build products that benefit diverse groups.
Distributed across a number of Host Institutions, our staff team supports the mission and operations of GA4GH.
Curious who we are? Meet the people and organisations across six continents who make up GA4GH.
More than 500 organisations connected to genomics — in healthcare, research, patient advocacy, industry, and beyond — have signed onto the mission and vision of GA4GH as Organisational Members.
These core Organisational Members are genomic data initiatives that have committed resources to guide GA4GH work and pilot our products.
This subset of Organisational Members whose networks or infrastructure align with GA4GH priorities has made a long-term commitment to engaging with our community.
Local and national organisations assign experts to spend at least 30% of their time building GA4GH products.
Anyone working in genomics and related fields is invited to participate in our inclusive community by creating and using new products.
Wondering what GA4GH does? Learn how we find and overcome challenges to expanding responsible genomic data use for the benefit of human health.
Study Groups define needs. Participants survey the landscape of the genomics and health community and determine whether GA4GH can help.
Work Streams create products. Community members join together to develop technical standards, policy frameworks, and policy tools that overcome hurdles to international genomic data use.
GIF solves problems. Organisations in the forum pilot GA4GH products in real-world situations. Along the way, they troubleshoot products, suggest updates, and flag additional needs.
GIF Projects are community-led initiatives that put GA4GH products into practice in real-world scenarios.
The GIF AMA programme produces events and resources to address implementation questions and challenges.
NIF finds challenges and opportunities in genomics at a global scale. National programmes meet to share best practices, avoid incompatabilities, and help translate genomics into benefits for human health.
Communities of Interest find challenges and opportunities in areas such as rare disease, cancer, and infectious disease. Participants pinpoint real-world problems that would benefit from broad data use.
The Technical Alignment Subcommittee (TASC) supports harmonisation, interoperability, and technical alignment across GA4GH products.
Find out what’s happening with up to the minute meeting schedules for the GA4GH community.
See all our products — always free and open-source. Do you work on cloud genomics, data discovery, user access, data security or regulatory policy and ethics? Need to represent genomic, phenotypic, or clinical data? We’ve got a solution for you.
All GA4GH standards, frameworks, and tools follow the Product Development and Approval Process before being officially adopted.
Learn how other organisations have implemented GA4GH products to solve real-world problems.
Help us transform the future of genomic data use! See how GA4GH can benefit you — whether you’re using our products, writing our standards, subscribing to a newsletter, or more.
Join our community! Explore opportunities to participate in or lead GA4GH activities.
Help create new global standards and frameworks for responsible genomic data use.
Align your organisation with the GA4GH mission and vision.
Want to advance both your career and responsible genomic data sharing at the same time? See our open leadership opportunities.
Join our international team and help us advance genomic data use for the benefit of human health.
Discover current opportunities to engage with GA4GH. Share feedback on our products, apply for volunteer leadership roles, and contribute your expertise to shape the future of genomic data sharing.
Solve real problems by aligning your organisation with the world’s genomics standards. We offer software dvelopers both customisable and out-of-the-box solutions to help you get started.
Learn more about upcoming GA4GH events. See reports and recordings from our past events.
Speak directly to the global genomics and health community while supporting GA4GH strategy.
Be the first to hear about the latest GA4GH products, upcoming meetings, new initiatives, and more.
Questions? We would love to hear from you.
Read news, stories, and insights from the forefront of genomic and clinical data use.
Attend an upcoming GA4GH event, or view meeting reports from past events.
See new projects, updates, and calls for support from the Work Streams.
Read academic papers coauthored by GA4GH contributors.
Listen to our podcast OmicsXchange, featuring discussions from leaders in the world of genomics, health, and data sharing.
Check out our videos, then subscribe to our YouTube channel for more content.
View the latest GA4GH updates, Genomics and Health News, Implementation Notes, GDPR Briefs, and more.
Discover all things GA4GH: explore our news, events, videos, podcasts, announcements, publications, and newsletters.
5 Nov 2018
Nearly six years ago, 50 colleagues came together in a conference room in New York City to discuss the future of genomics. With delegates from eight different countries, they represented an international, interdisciplinary field of researchers, clinicians, and professionals who knew that massive amounts of data were on the horizon and that a plan was needed to make the most of it all. One thing was clear: the plan needed to be global.

Dear Colleagues,
Nearly six years ago, 50 colleagues came together in a conference room in New York City to discuss the future of genomics. With delegates from eight different countries, they represented an international, interdisciplinary field of researchers, clinicians, and professionals who knew that massive amounts of data were on the horizon and that a plan was needed to make the most of it all.
One thing was clear: the plan needed to be global. Any solution we developed had to be relevant to all nations and cultures.
The outcome of that meeting was the Global Alliance for Genomics and Health (GA4GH). Since then, GA4GH has made great strides. We’ve grown into a legitimate international standards body that is setting the course for large-scale genomic data sharing. We are maintaining the key file formats used by the international community and are rolling out a series of APIs and specifications that any nation can theoretically take up into their existing workflows to make their data interoperable with the world’s data.
In practice, however, we are quite aware that much more work is needed to achieve the first G, “Global”, in GA4GH.
We are now putting real attention toward changing that. The recent plenary meeting held in Basel, Switzerland attracted a record 425+ individuals from 25 countries. We had speakers from 12 countries and heard from national genomic data initiatives underway in nine regions (Switzerland, Finland, England, France, the United States, Australia, Japan, and across the continents of both Africa and Europe). We had a great workshop among national initiatives for sharing resources and learnings as well as instructions on how to get involved with the GA4GH Work Streams.
But most importantly, we announced an open call for new GA4GH Driver Projects, where the key focus will be on increasing our global representation. We currently have 15 of these leading genomic data initiatives “driving” the development activities of our Work Streams, helping us build tools and pilot them in real world scenarios. Collectively, they are headquartered in five separate countries, a map of which I presented in Basel:
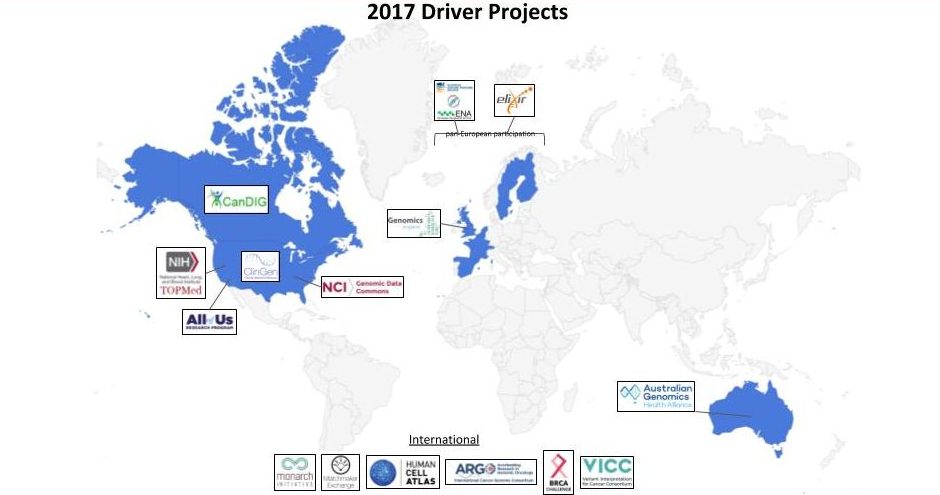
As you can see, there’s still a lot of gray. We’re hoping to turn some of those gray bits blue in this next group of Driver Projects.
In that same presentation, I showed another map that tracks countries with a national medical genome initiative as well as countries with large scale national cohorts (>100K participants):
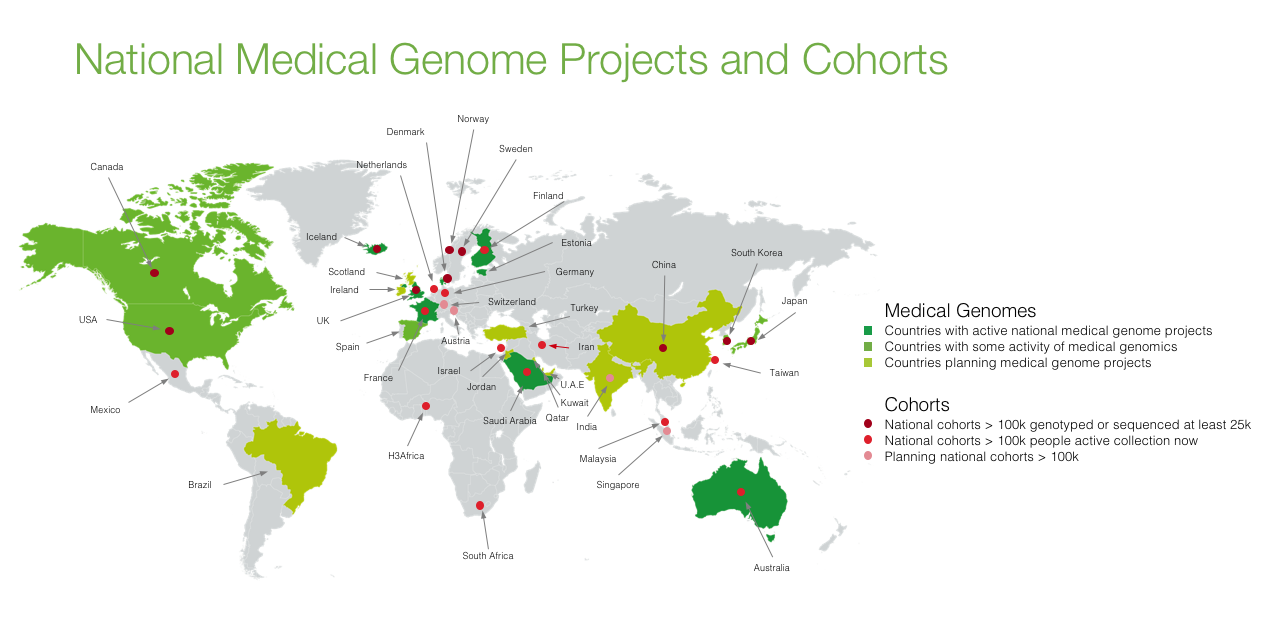
Ultimately, we’re hoping the two maps will start to look more and more alike; that is to say, we hope for GA4GH to truly be driven by the entire global community engaged in medical genomics. (By the way, please help us ensure the green map is correct — if you don’t see your project represented above, please let us know.)
Please review the criteria for becoming a GA4GH Driver Project and submit your application by November 30. If you don’t think your project is a good fit as a Driver project, don’t let that stop you from getting involved. Find the Work Stream that most aligns with the standards your team needs and reach out to the relevant manager to find out how you can participate. Take a look at our toolkit of frameworks and specifications (such as the recently released Beacon, WES, and refget APIs) and reach out to a Work Stream Manager to get support with your implementation.
There are 193 countries in the UN. I believe that all of them will integrate genomics into their national health systems in some way in the next decade. If our work is to impact health outcomes for patients around the world, GA4GH needs to tap into the knowledge across that entire geographic spread. We need all your help to succeed in this ambition.
Notable GA4GH Publications since July:
